Since its inception in September, the facility has been running at near full capacity with various pilot projects to test the system and familiarize the staff with its operation.
Facility staff worked with researchers from three different departments across campus so far, including Institute of Biological Chemistry (IBC), Plant Pathology, and School of Biological Sciences.

To date, researchers were particularly interested in using our Walz Pulse Amplitude Modulation (PAM) camera to measure chlorophyll fluorescence to determine photosynthetic efficiency in the context of various stress conditions. For example, postdoc Meng Li along with PI Helmut Kirchhoff of IBC were able to measure and process multiple fluorescence measurements of 41 plants a day, including protocols at night when the plants are naturally in a dark-adapted state to measure the maximum fluorescence and photosynthetic efficiency. This is particularly useful because it sets the baseline from which to compare daytime photosynthetic efficiency against. Plants naturally decrease their efficiency with higher light exposure to protect themselves from too much energy. Historically, capturing robust maximum fluorescence meant either staying up all night or disrupting the natural light rhythm for the plants. Li and Kirchhoff are interested in ion fluxes across the thylakoid membrane of the chloroplast under rapid and unpredictable changes in light intensity and how well their mutants can handle these conditions.
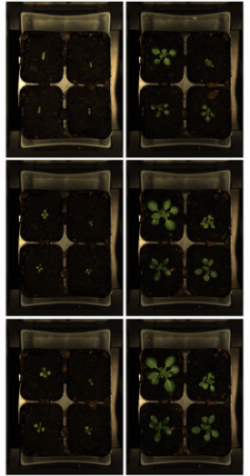
Graduate student Samantha The and PI Mechthild Tegeder, from the School of Biological Sciences, are looking at the effect of different stress conditions on amino acid partitioning in Arabidopsis. In preliminary analyses we can detect differential growth among different mutants under drought and salt stress conditions. This project highlights the advantage of the new facility – the ability to quickly and efficiently screen 168 plants for important phenotypic differences with minimal user interaction to identify the important mutations. The and Tegeder hope the data from this pilot study will help them refine their hypotheses and win funding from NSF for a larger study.
